-Search query
-Search result
Showing all 21 items for (author: isupov & mn)

EMDB-19608: 
Sulfolobus acidocaldarius threads (0406) filament.
Method: helical / : Isupov MN, Gaines M, McLaren M, Daum B

EMDB-18119: 
Sulfolobus acidocaldarius AAP filament.
Method: helical / : Isupov MN, Gaines M, Daum B, McLaren M

EMDB-18127: 
S-layer of archaeon Sulfolobus acidocaldarius by subtomogram averaging
Method: subtomogram averaging / : Gambelli L, McLaren MJ, Daum B

EMDB-17369: 
A CHIMERA construct containing human SARM1 ARM and SAM domains and C. elegans TIR domain.
Method: single particle / : Isupov MN, Opatowsky Y

EMDB-17370: 
C. elegans TIR-1 protein.
Method: single particle / : Isupov MN, Opatowsky Y

EMDB-15530: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 10.0
Method: single particle / : Gambelli L, Isupov MN, Daum B

EMDB-15531: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 7.0
Method: single particle / : Gambelli L, Isupov MN, Daum B

EMDB-14778: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the half-closed conformation
Method: single particle / : Freda I, Montemiglio LC, Tramonti A, Contestabile R, Vallone B, Savino C, Exertier C, Bolognesi M, Chaves Sanjuan A

EMDB-14801: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C2 symmetry
Method: single particle / : Freda I, Montemiglio LC, Tramonti A, Contestabile R, Vallone B, Exertier C, Savino C, Chaves Sanjuan A, Bolognesi M

EMDB-14852: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the closed conformation, C1 symmetry
Method: single particle / : Freda I, Montemiglio LC, Tramonti A, Contestabile R, Vallone B, Exertier C, Savino C, Chaves Sanjuan A, Bolognesi M

EMDB-14960: 
Cryo-EM structure of holo-PdxR from Bacillus clausii bound to its target DNA in the open conformation
Method: single particle / : Freda I, Montemiglio LC, Tramonti A, Contestabile R, Vallone B, Exertier C, Savino C, Chaves Sanjuan A, Bolognesi M

EMDB-17448: 
Spraguea lophii ribosome in the closed conformation by cryo sub tomogram averaging
Method: subtomogram averaging / : Gil Diez P, McLaren M, Isupov MN, Daum B, Conners R, Williams B

EMDB-17457: 
Spraguea lophii ribosome dimer
Method: subtomogram averaging / : Gil Diez P, McLaren M, Isupov MN, Daum B, Conners R, Williams B

EMDB-14635: 
S-layer protein SlaA from Sulfolobus acidocaldarius at pH 4.0
Method: single particle / : Gambelli L, Isupov MN, Daum B

EMDB-13892: 
Spraguea lophii ribosome
Method: single particle / : Gil Diez P, McLaren M

EMDB-13546: 
Sulfolobus acidocaldarius 0406 filament.
Method: helical / : Isupov MN, Gaines M, Daum B

EMDB-12875: 
The archaellum of Methanocaldococcus villosus
Method: helical / : Isupov M, Gambelli L

EMDB-11187: 
hSARM1 GraFix-ed
Method: single particle / : Sporny M, Guez-Haddad J, Khazma T, Yaron A, Dessau M, Mim C, Isupov MN, Zalk R, Hons M, Opatowsky Y
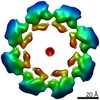
EMDB-11190: 
SARM1 SAM1-2 domains
Method: single particle / : Sporny M, Guez-Haddad J, Khazma T, Yaron A, Dessau M, Mim C, Isupov MN, Zalk R, Hons M, Opatowsky Y

EMDB-11191: 
SARM1 SAM1-2 domains
Method: single particle / : Sporny M, Guez-Haddad J, Khazma T, Yaron A, Dessau M, Mim C, Isupov MN, Zalk R, Hons M, Opatowsky Y

EMDB-11834: 
hSARM1 NAD+ complex
Method: single particle / : Sporny M, Guez-Haddad J
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
